Great tip
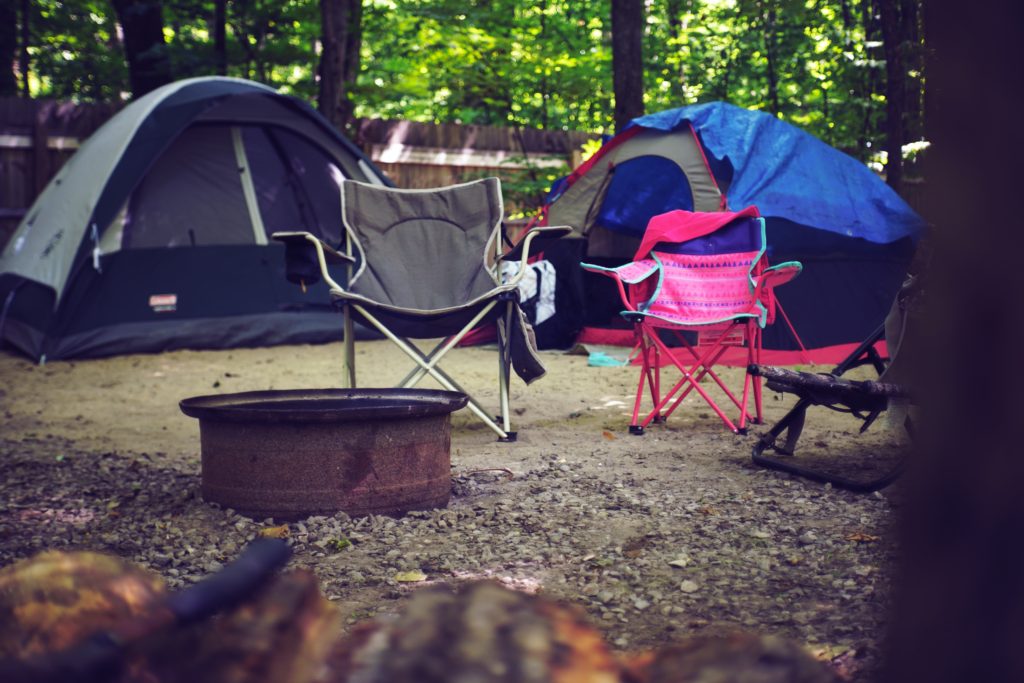
Another great tip is to bring some newspaper with you as it will help dry out your tent, gear and anything else that gets wet. You can stick a few rolled-up clumps of it into your shoes at night to speed up the drying process and also helps with fire starting in a wet environment! For more detail about overnatting kristiansand.
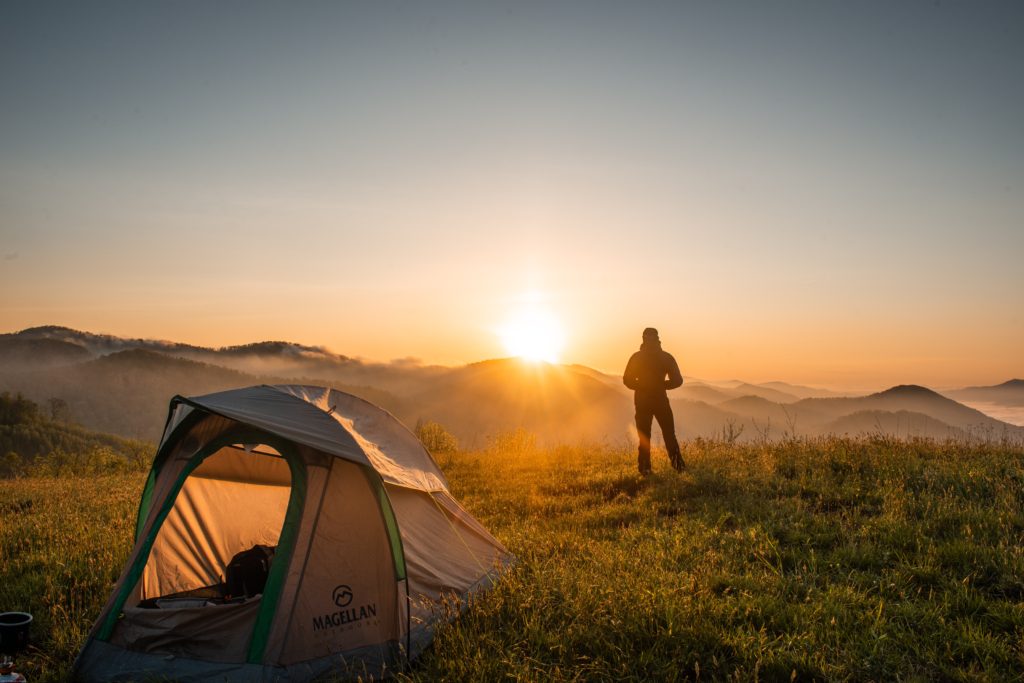
A good rain coat and poncho will help keep you dry while hiking, but it’s also important to have enough layers to stay warm in case of extended periods of rainy weather. It important to know that weather can change quickly, especially in the fall. Always check the forecast for your camping destination to best prepare for your trip.